Genetic data
Whole genome sequencing
UK Biobank's whole genome sequencing data on all 500,000 participants - the biggest whole genome dataset in the world - is now available to approved researchers on the UK Biobank Research Analysis Platform.
It will transform the way in which scientists study the genetic determinants of a wide range of health outcomes, providing information that will complement and enhance the existing genotyping and exome data.
The Medical Research Council provided funding to UK Biobank in 2018 for a pilot project (the Vanguard) to perform whole-genome sequencing on 50,000 participants, which was undertaken by the Wellcome Sanger Institute, Cambridge.
A consortium of government, industry and charity then came together to fund whole genome sequencing of the remaining 450,000 participants. This project was funded by:
- UK Government’s research and innovation agency, UK Research and Innovation (UKRI), through the Industrial Strategy Challenge Fund
- The Wellcome Trust
- A consortium of industry partners: Amgen, AstraZeneca, GlaxoSmithKline and Johnson & Johnson.
Decode Genetics and the Wellcome Sanger Institute carried out the sequencing using Illumina Novaseq technology.
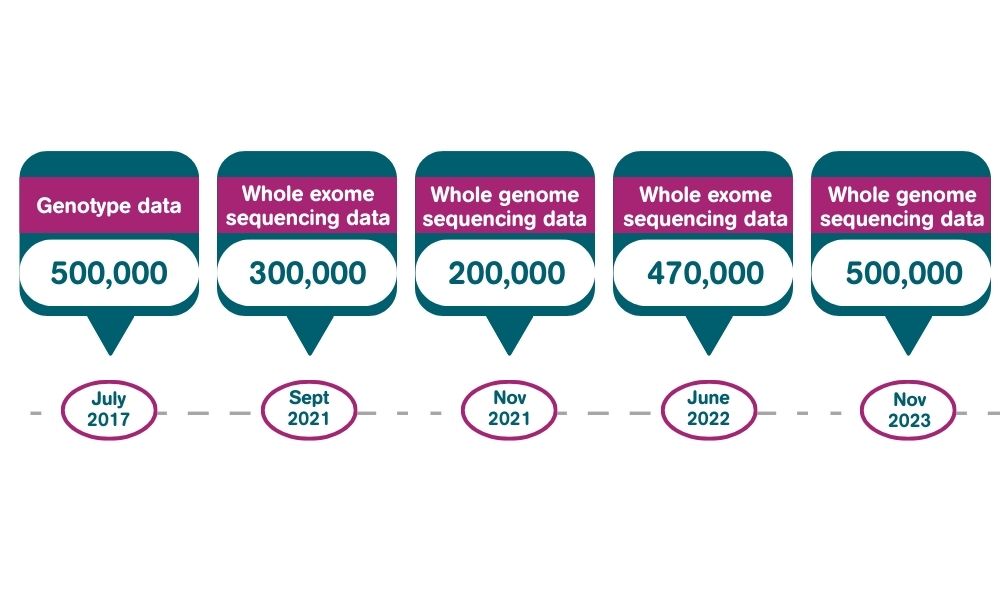
Data for 200,000 genomes was released in 2021.
Data for 500,000 whole genomes is now available to approved researchers in the UK Biobank Research Analysis Platform.
Whole genome sequencing data access
The data is available only to researchers who have been approved by UK Biobank and are using the UK Biobank Research Analysis Platform (UKB-RAP). To use the UKB-RAP you must register with UK Biobank, apply for approval for data access and then sign up to the UKB-RAP itself. You can browse the summary statistics for all UK Biobank participants free of charge using the Allele Frequency Browser.
Accessing genetic data
Please note that some information regarding data access and download in the below FAQs may be out of date. For instructions on accessing or downloading genetic data, please refer to our Data Access Guide.

Genome-wide genotyping
Genome-wide genotyping was performed on all UK Biobank participants using the UK Biobank Axiom Array. Approximately 850,000 variants were directly measured, with > 90million variants imputed using the Haplotype Reference Consortium and UK10K + 1000 Genomes reference panels.
You can view the full Axiom array SNP list by downloading the csv file*. Alternatively, you can use the Genomic Search facility to find specific genetic loci of interest that are measured on the array. Imputed data using different reference panels are planned to be made available in the future.
Genotyping and Imputation FAQs
Related publication

Whole exome sequencing
Whole-exome sequencing measures the regions of the genome (about 2%) that are involved in coding for proteins and is particularly suitable for identifying disease-causing and/or rare genetic variants.
A vanguard exome sequencing project on the first 50,000 participants was performed by Regeneron and GlaxoSmithKline. A further consortium (comprising Regeneron, AbbVie, Alnylam Pharmaceuticals, AstraZeneca, Biogen, Pfizer, Takeda and Bristol-Myers Squibb) have completed the exome sequencing project and data is available to researchers for 470,000 participants.
Exome sequencing FAQs
Related publication
Whole exome sequencing and characterization of coding variation in 49,960 individuals in the UK Biobank
Cristopher V. Van Hout et alExplore our other data
Last updated